Temperature sensing in plants
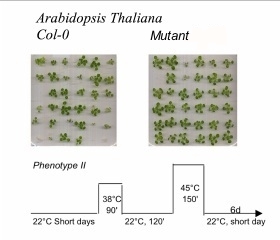
Land plants need to rapidly detect mild temperature increments at physiological temperatures to establish thermotolerance against the damages of an upcoming heat stress. The onset of plant thermotolerance requires the early perception of an upcoming heat stress by timely responding by accumulating heat shock proteins (Hsps), many of which, molecular chaperones that can protect proteins and membranes from heat-damage. Hence, land plants need precise molecular thermosensors, which have to be coupled to a specific signaling pathway to the cytoplasm and the nucleus to allow the upregulation of HSP genes. We found strong biochemical evidence for the existence of highly specific calcium channels in the plasma membrane of the moss Physcomitrella patens, which act as the most upstream heat-sensors of the plant cell (Saidi et al. 2009).
Our further aims are focused to establish the molecular identity of the Ca2+ channel serving as heat receptor(s) in moss P. patens and to characterize protein loss-of-function, especially with regards to the spectrum or intensity of chaperone expression and to the onset of acquired thermotolerance and correlate chaperone expression with Ca2+ transients. Using moss as a model system we would like to identify the equivalent heat receptor genes in higher plants and to characterize their phenotypes in terms of HSP expression and acquired thermotolerance.

Molecular chaperones as nanomachines
By virtue of their general ability to bind (hold) translocating or unfolding polypeptides otherwise doomed to aggregate, molecular chaperones are commonly dubbed “holdases”. Yet, chaperones also carry physiological functions that do not necessitate prevention of aggregation, such as altering the native states of proteins, as in the disassembly of SNARE complexes and clathrin coats. To carry such physiological functions, major members of the Hsp70, Hsp110, Hsp100, and Hsp60/CCT chaperone families act as catalytic unfolding enzymes or unfoldases that drive iterative cycles of protein binding, unfolding/pulling, and release. One unfoldase chaperone may thus successively convert many misfolded or alternatively folded polypeptide substrates into transiently unfolded intermediates, which, once released, can spontaneously refold into low-affinity native products. Whereas during stress, a large excess of non-catalytic chaperones in holding mode may optimally prevent protein aggregation, after the stress, catalytic disaggregases and unfoldases may act as nanomachines that use the energy of ATP hydrolysis to repair proteins with compromised conformations. Thus, holding and catalytic unfolding chaperones can act as primary cellular defenses against the formation of early misfolded and aggregated proteotoxic conformers in order to avert or retard the onset of degenerative protein conformational diseases.
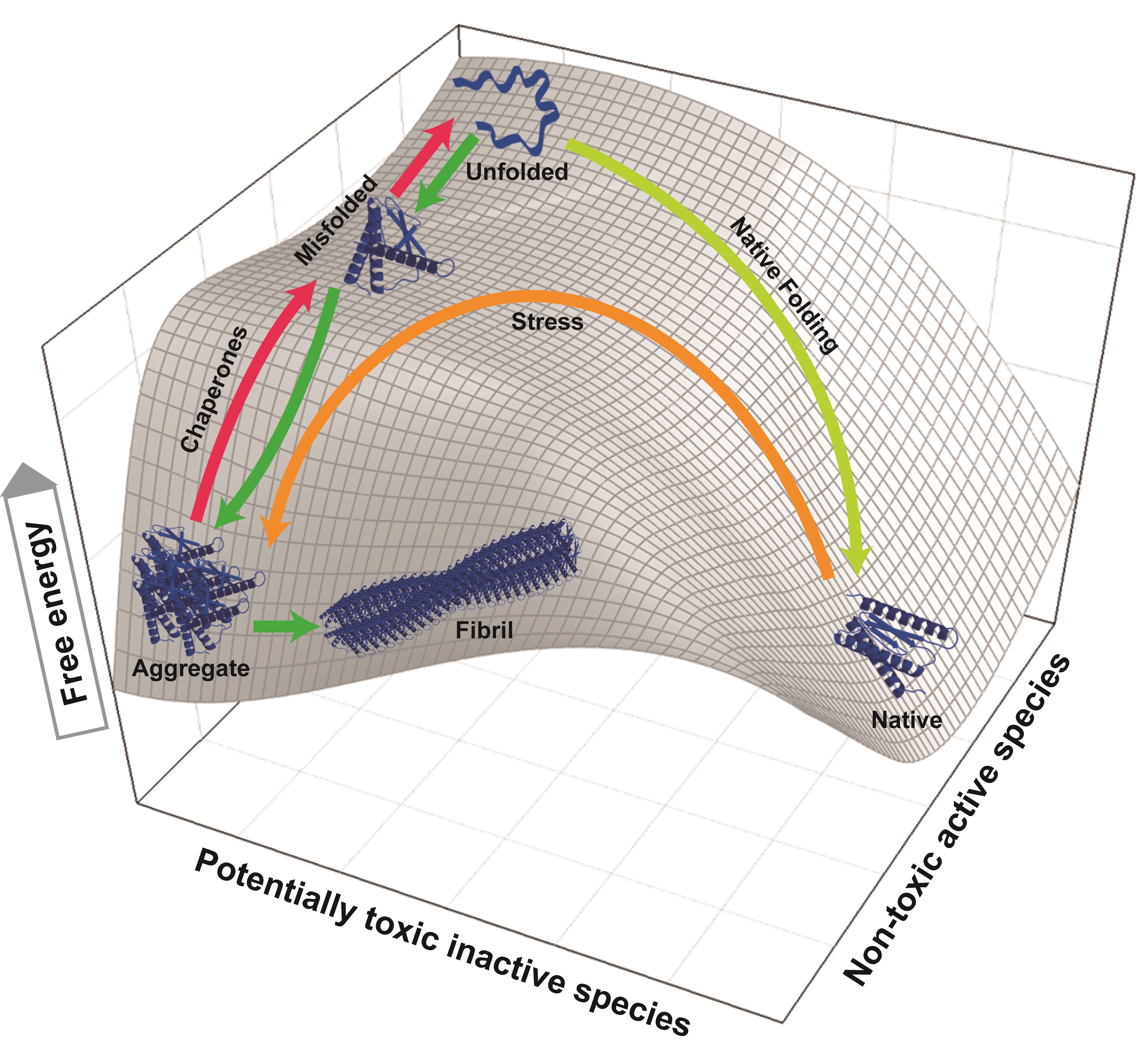
Disaggregating and unfolding chaperones transiently increase the free energy of misfolded or alternatively folded substrates but not of natively folded products. The 3D mesh-plot shows a typical unfolded polypeptide with the highest free energy, which can spontaneously reach to lower free energy states, either by folding to the native conformation (right) or by misfolding to the aggregated state (left). When conditions are not favorable for native folding, the unfolded polypeptide may prefer undertaking the misfolding pathway to aggregation (green arrows). Native proteins under stress gaining free energy may partially unfold to a state from which it can readily seek a more stable misfolded state [3] and further aggregate (brown arrow) [2]. ATP-fueled unfoldase chaperones drive the substrate uphill the free energy barrier (red arrows) by converting stable aggregates and misfolded species into unfolded products with a higher free energy. From there, if conditions are favorable to the end product, the unfolded species can collapse to the stable native state.
Quantitative proteomics of chaperones systems in cultured cells
In the crowded environment of human cells, folding of nascent polypeptides and refolding of stress-unfolded proteins is error prone. Accumulation of cytotoxic misfolded and aggregated species may cause cell death, tissue loss, degenerative conformational diseases, and aging. Nevertheless, young cells effectively express a network of molecular chaperones and folding enzymes, termed here “the chaperome,” which can prevent formation of potentially harmful misfolded protein conformers and use the energy of adenosine triphosphate (ATP) to
rehabilitate already formed toxic aggregates into native functional proteins. In an attempt to extend knowledge of chaperome mechanisms in cellular proteostasis, we performed a meta-analysis of human chaperome using high-throughput proteomic data from 11 immortalized human cell lines. Chaperome polypeptides were about 10 % of total protein mass of human cells, half of which were Hsp90s and Hsp70s. Knowledge of cellular concentrations and ratios among chaperome polypeptides provided a novel basis to understand mechanisms by which the Hsp60, Hsp70, Hsp90, and small heat shock proteins (HSPs), in collaboration with cochaperones and folding enzymes, assist de novo protein folding, import polypeptides into organelles, unfold stress-destabilized toxic conformers, and control the conformal activity of native proteins in the crowded environment of the cell. Proteomic data also provided means to distinguish between stable components of chaperone core machineries and dynamic regulatory cochaperones.
.jpg)
Chaperones are among the most abundant proteins in HeLa cells. (a) Distributions of mean mass proportions and SD (in percent of total, upper graph), and copy number (lower graph) of each individual protein present in statistically significantly similar amounts per cell. Chaperome proteins are in black and non-chaperome proteins in gray. Means and SD are from three independent measurements from HeLa cells. (b) Distribution of mass proportions from total HeLa cell proteins of the main chaperome systems as indicated: Hsp90 with cochaperones, Hsp70 with cochaperones, Hsp60 with Hsp10 cochaperone, the small HSPs, and the foldases PPIs and PDIs, in the cytosol (white), the endoplasmic reticulum (ER, deep gray), and the mitochondria (Mito, light gray)(Geiger et al. 2012).
Functionality and mechanisms of Hsp70 chaperone system
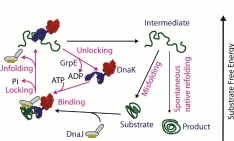
Stress, molecular crowding and mutations may jeopardize the native folding of proteins. Misfolded and aggregated proteins not only loose their biological activity, but may also disturb protein homeostasis, damage membranes and induce apoptosis. Molecular chaperones belong to several families of highly conserved proteins that favor the native folding of proteins, both as “holdases” that sequester hydrophobic regions in misfolding polypeptides and thus preventing protein aggregation, or as active unfoldases, that use the energy of ATP hydrolysis to unfold and disentangling misfolded and aggregated polypeptides. We try to understand central questions such as: By which mechanism may a single Hsp70 chaperone change the conformation of a stable misfolded polypeptide substrate? By which steps do Hsp70 and its co-chaperones use ATP to transforming inactive misfolded or alternatively folded polypeptides into active native proteins? Is chaperone-mediated protein refolding to the native state spontaneous or is it directed by the chaperone? Does Hsp70 behave as a true unfolding and/or folding catalyst capable of multiple turnovers to convert an excess of stable misfolded substrates into stable native products? What is the minimal ATP cost for transforming a single inactive misfolded polypeptide into an active, natively folded one? Chaperones are central cellular defenses against the formation of toxic protein conformers in protein-misfolding diseases, such as Alzheimer, Parkinson disease and aging in general.
In vitro reconstitution of human Hsp70 system and its implication in protein misfolding problems
There is plethora of literature indicating that the molecular chaperones and chaperone controlled proteases, such as the proteasome, belong to a cellular network that can prevent and reduce the formation of toxic aggregates and possibly eliminate already formed toxic protein aggregates in neurodegenerative diseases. The respective analogues of Hsp70 system in bacteria are Dnak/DnaJ/GrpE. 70-kDa heat shock proteins (Hsp70s) assist a wide range of folding processes, including the folding and assembly of newly synthesized proteins, refolding of misfolded and aggregated proteins, membrane translocation of organellar and secretory proteins, and control of the activity of regulatory proteins. My main objective is to develop efficient in vitro human Hsp70 chaperone system and its co-chaperones Hsp40/NEF. The in vitro bacterial Hsp70 machinery is already well established in the Goloubinoff Laboratory and this will serve as control for my work. I shall primarily study the kinetic parameters and energy cost of the human Hsp70 machinery. I will also focus on the possible implication of the Hsp70 machinery in protein conformation diseases.



.jpg)
